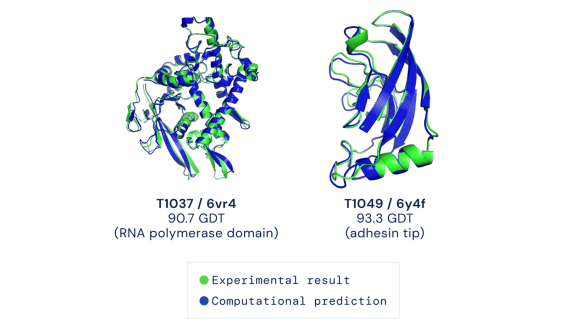
DeepMind, a company based in London and owned by Google, announced that it had predicted the three-dimensional structures of more than 200 million proteins using AlphaFold.
This is the entire protein universe known to scientists today.
What is AlphaFold?
- AlphaFold is an AI-based protein structure prediction tool.
- It is based on a computer system called deep neural network.
- Inspired by the human brain, neural networks use a large amount of input data and provide the desired output exactly like how a human brain would.
- The real work is done by the black box between the input and the output layers, called the hidden networks. AlphaFold is fed with protein sequences as input.
- When protein sequences enter through one end, the predicted three-dimensional structures come out through the other.
- It is like a magician pulling a rabbit out of a hat.
How does AlphaFold work?
- It uses processes based on “training, learning, retraining and relearning.”
- The first step uses the available structures of 1,70,000 proteins in the Protein Data Bank (PDB) to train the computer model.
- Then, it uses the results of that training to learn the structural predictions of proteins not in the PDB.
- Once that is done, it uses the high-accuracy predictions from the first step to retrain and relearn to gain higher accuracy of the earlier predictions.
- By using this method, AlphaFold has now predicted the structures of the entire 214 million unique protein sequences deposited in the Universal Protein Resource (UniProt)
What are the implications of this development?

- Proteins are the business ends of biology, meaning proteins carry out all the functions inside a living cell.
- Therefore, knowing protein structure and function is essential to understanding human diseases.
- Scientists predict protein structures using x-ray crystallography, nuclear magnetic resonance spectroscopy, or cryogenic electron microscopy.
- These techniques are not just time-consuming, they often take years and are based mainly on trial-and-error methods.
- The development of AlphaFold changes all of that.
- It is a watershed movement in science and structural biology in particular.
What does this development mean for India?
- Vaccine development: Understanding the accurate structures of COVID-19 virus proteins in days rather than years will accelerate vaccine and drug development against the virus.
- Structural biology: From the seminal contribution of G. N. Ramachandran in understanding protein structures to the present day, India is no stranger to the field and has produced some fine structural biologists.
Back2Basics: Proteins
- Protein is found throughout the body—in muscle, bone, skin, hair, and virtually every other body part or tissue.
- It makes up the enzymes that power many chemical reactions and the hemoglobin that carries oxygen in your blood.
- At least 10,000 different proteins make you what you are and keep you that way.
- Protein is made from twenty-plus basic building blocks called amino acids.
- Because we don’t store amino acids, our bodies make them in two different ways: either from scratch or by modifying others.
- Nine amino acids—histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine—known as the essential amino acids, must come from food.
- Chemically, amino acids are organic compounds made of carbon, hydrogen, nitrogen, oxygen or sulfur.
- There are seven types of proteins: antibodies, contractile proteins, enzymes, hormonal proteins, structural proteins, storage proteins, and transport proteins.
UPSC 2022 countdown has begun! Get your personal guidance plan now! (Click here)